I’ve been having a lot of fun over the last 18 months re-discovering my guitar, playing, and singing… I’ve even been writing songs & making recordings, some of which I’ve posted on my soundcloud account. A few of these songs formed the basis for a live performance I gave at Oxford University in November 2024, at an event entitled “The Museum of Consciousness” curated by Karl H Smith.
Author Archives: David R Glowacki
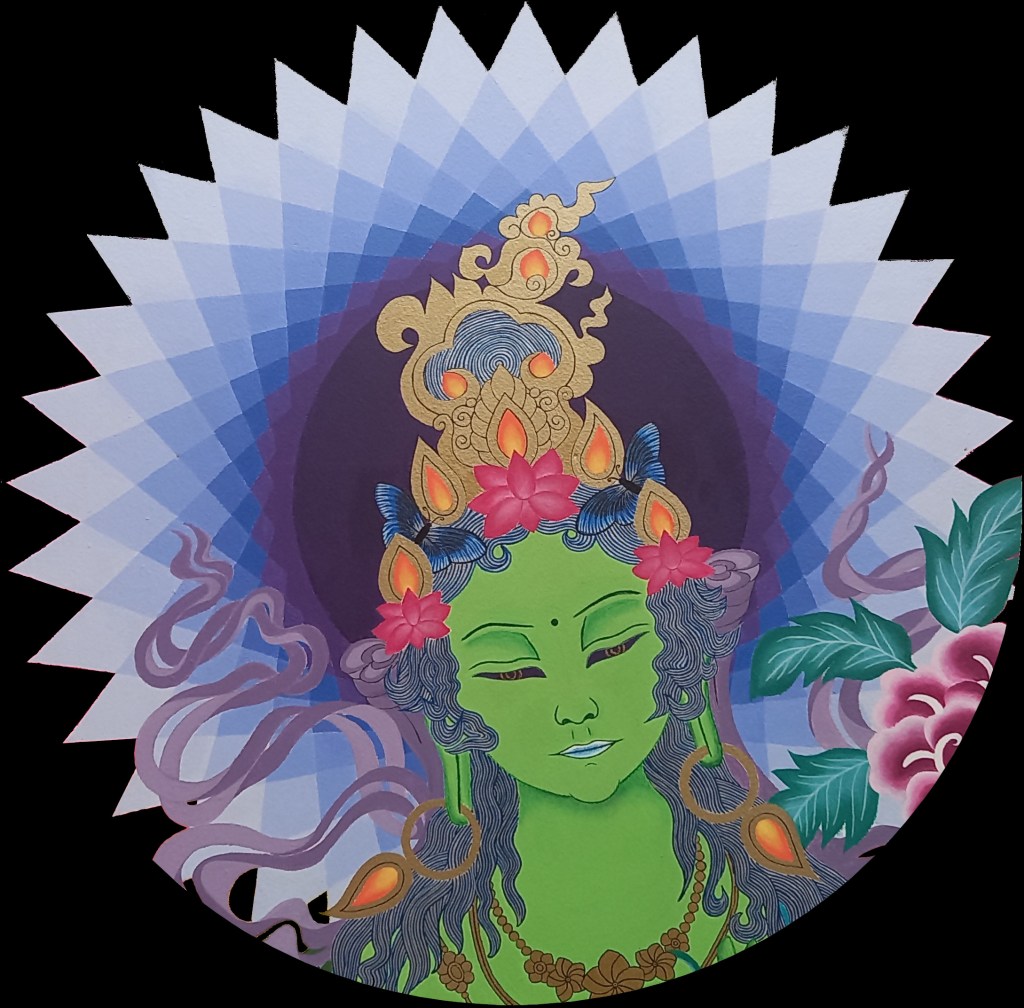
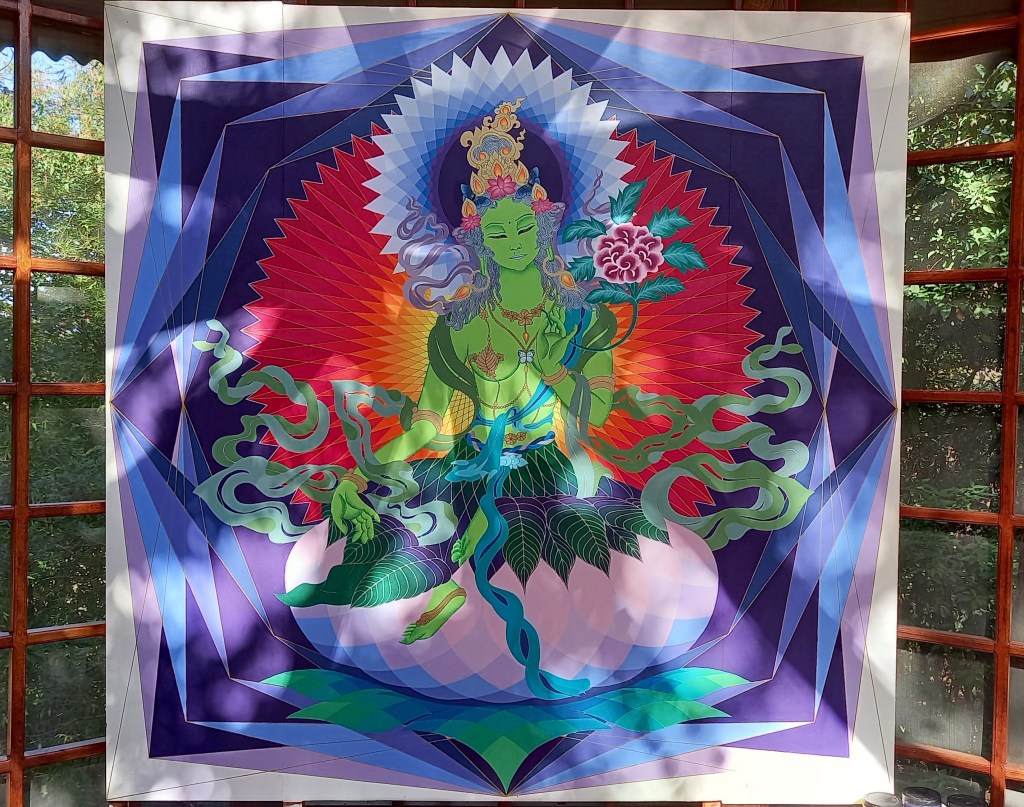
Green Tara in Jaén, Perú
Ripple.
I’m writing a paper about my near-death experience, which is what has underpinned a lot of my aesthetic over these last years. This video illustrates how I perceived my body – as a pulsing light. The light intensity grew with each inhale, and diminished with each exhale. Overall, the amplitude envelope was decreasing. I knew that at the moment it was extinguished, I would begin resting. In another dimension. It seems it wasn’t the time.
Entropy
Here’s a demo video from ‘Gaussian Fields’, a new physics art C++ real-time simulation code I have been working on. It’s a generative framework, so you never know exactly what patterns will emerge once you set the dynamics in motion. It’s cool to see intricate and varied complexity evolve from such simple initial conditions….
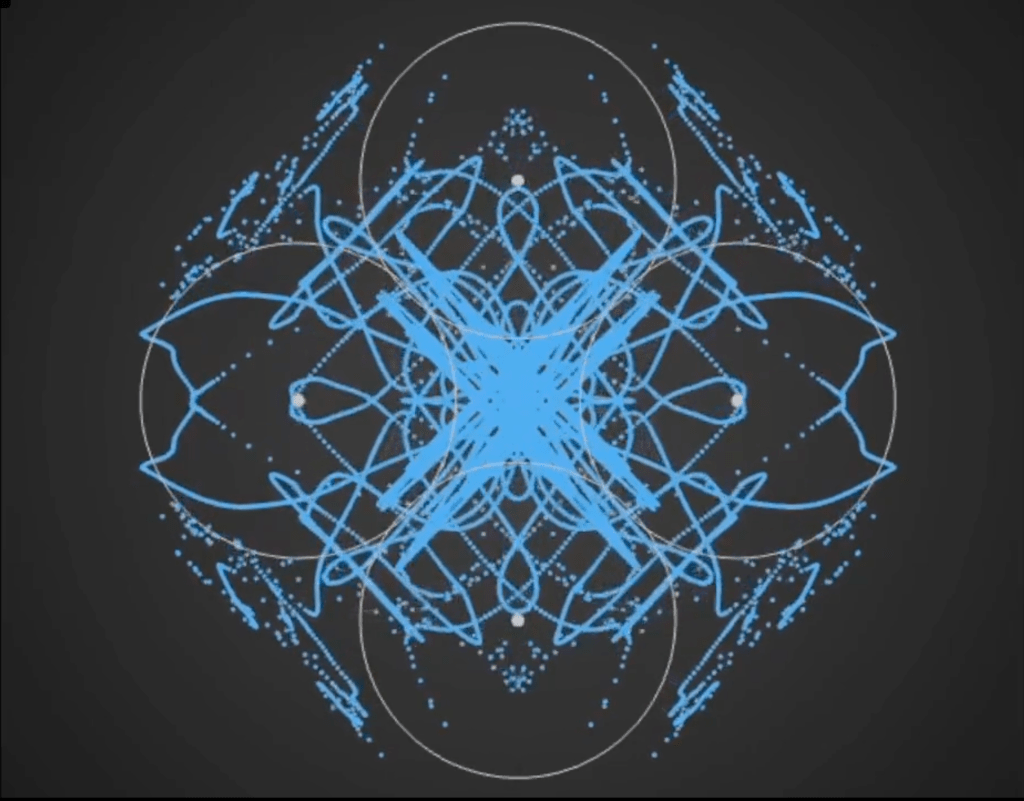
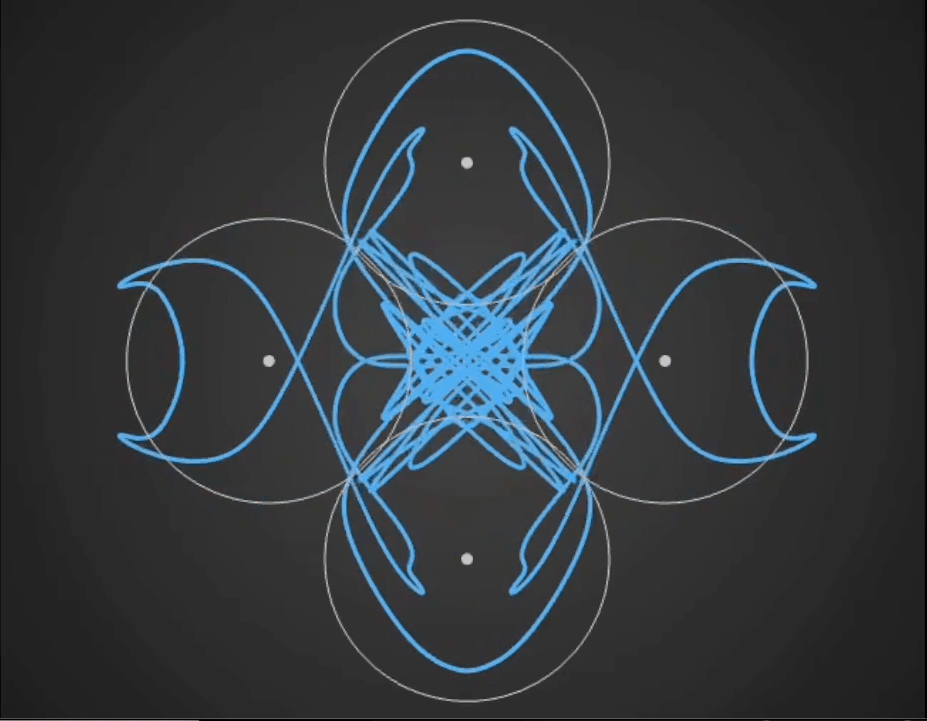
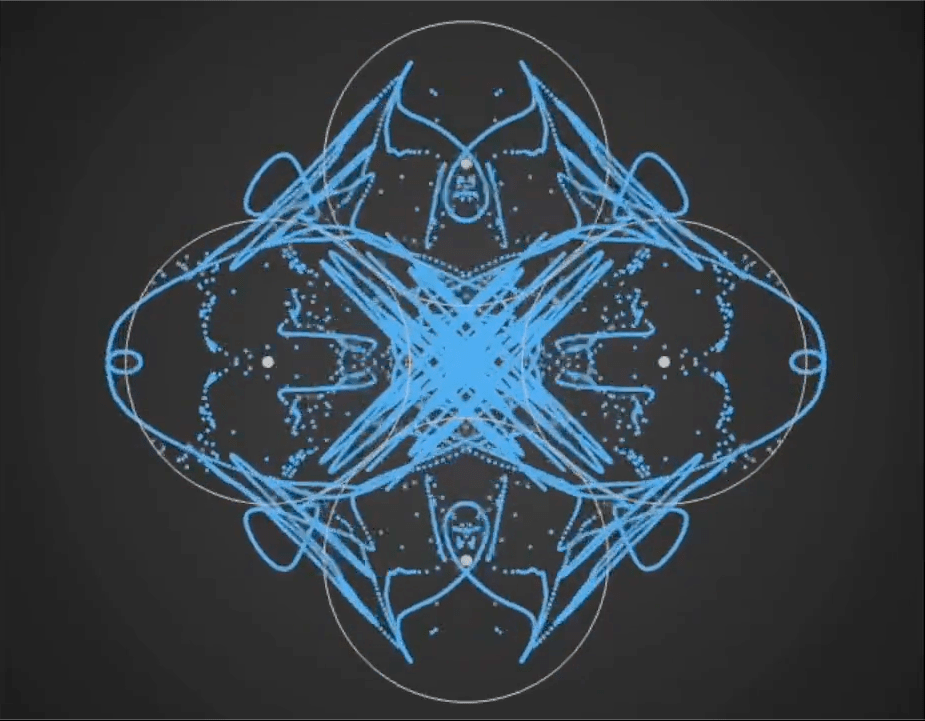
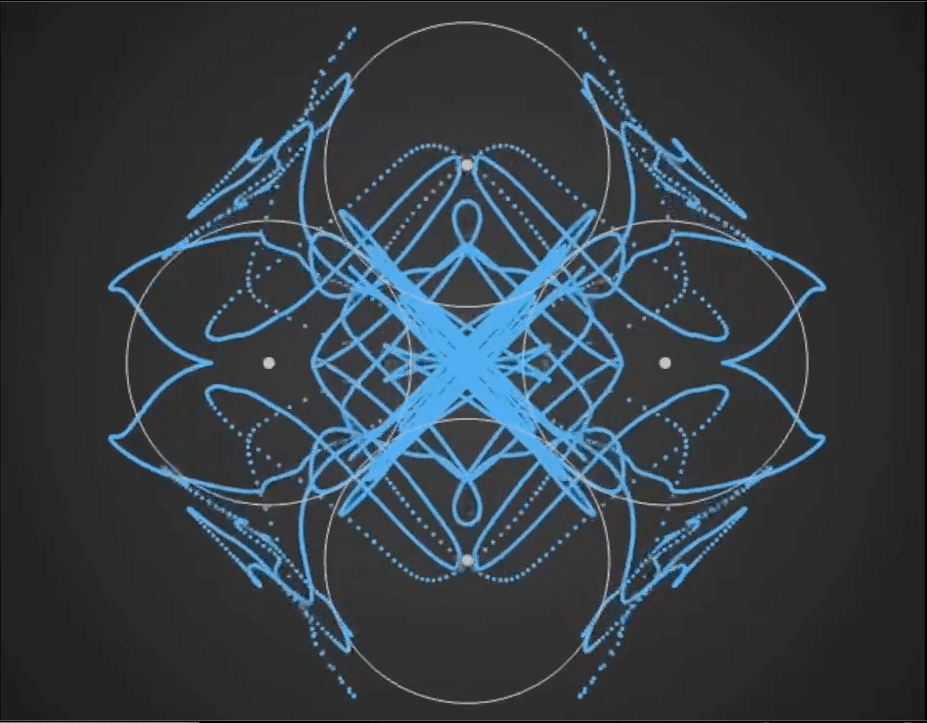


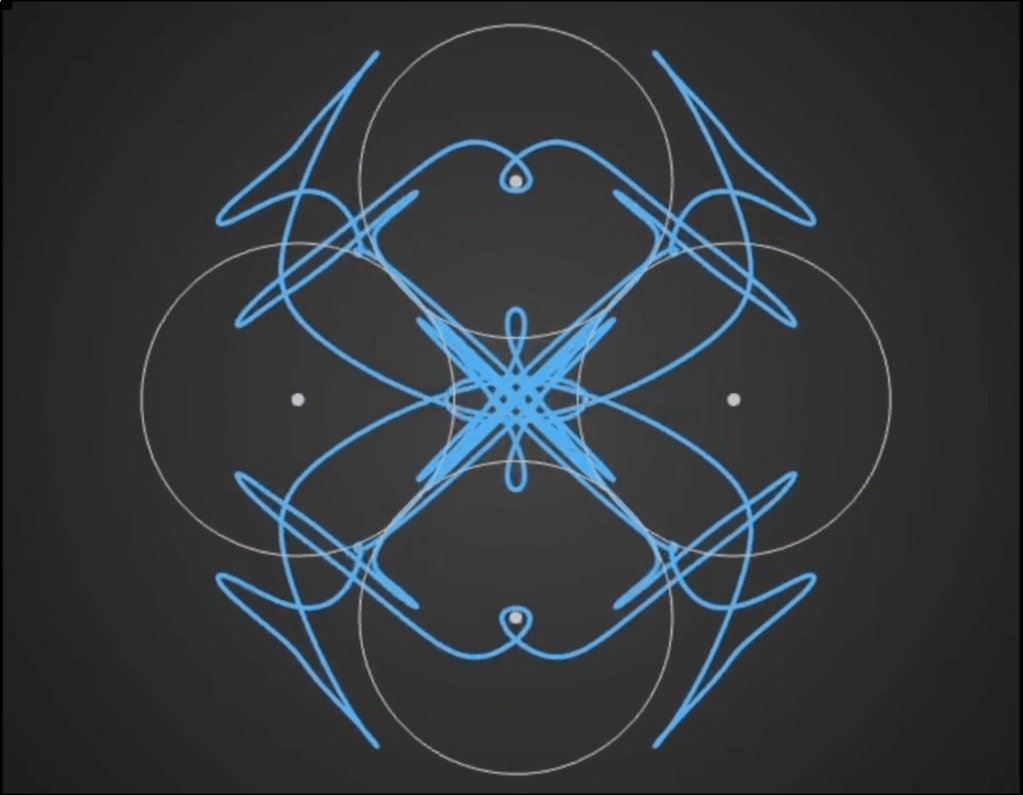
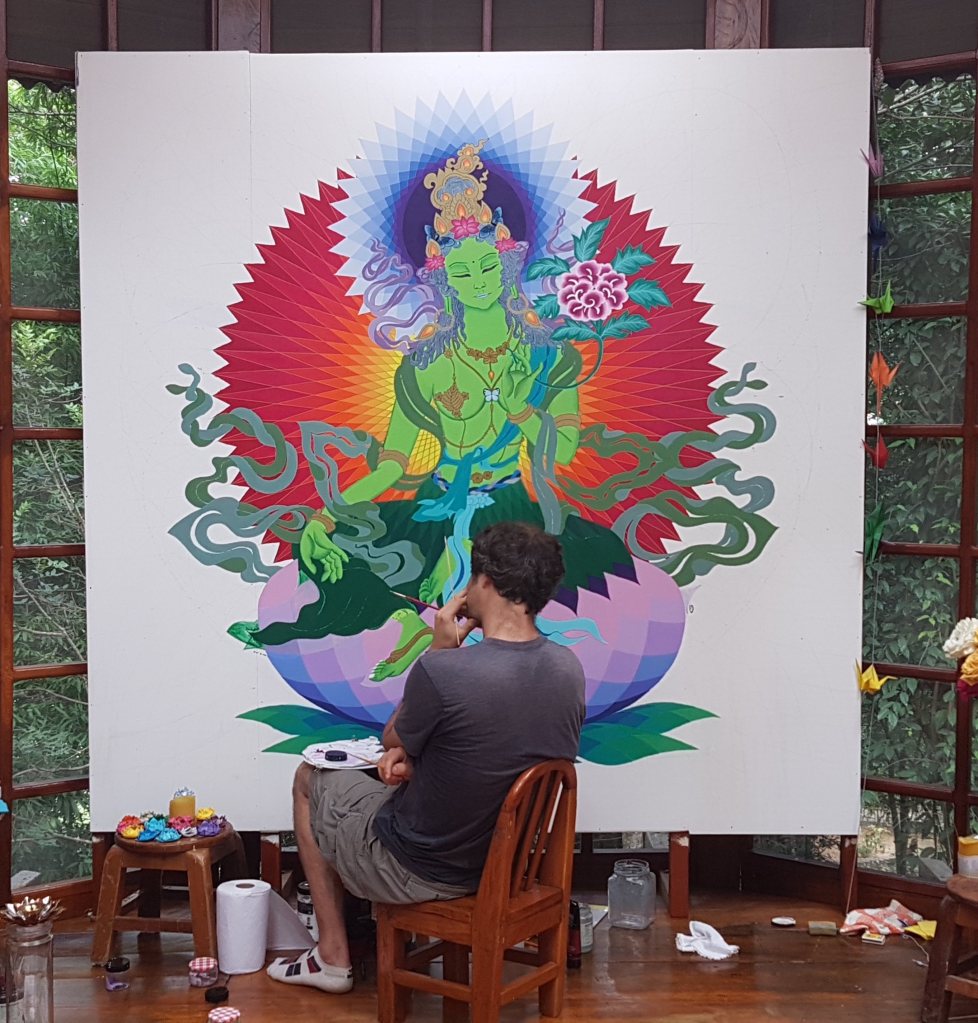
Tara in progress…
I have been carrying out an intensive aesthetic research pilot at the home of Maria Cristina Mendoza Vidal (an ayahuascera near Jaén, Peru) exploring how so-called ‘deity yoga’ practice (from Vajrayana buddhism) can be adapted for those in psychedelic altered states who have no prior deity yoga experience.
As part of this research, I have spent many hours engaged in eyes-open Tārā deity yoga practice in a jungle-hut-turned-multimedia-art-studio in Perú creating a Tārā mural whilst communing with the native plants ayahuasca and chakruna. The details of the aesthetic theory which has arisen during this process, and which has guided Tārā’s emergence, are outlined in a recent research article. This Tārā has dark eyes. More soon.

IRL 2.0
Intangible Realities Laboratory (IRL) headquarters has moved to the CiTIUS | Centro Singular de Investigación en Tecnoloxías Intelixentes in Santiago de Compostela. To celebrate the incarnation of IRL 2.0, we have a shiny new website!
www.intangiblerealitieslab.org

Citizen science studies comparing group VR to psychedelics

To read more, check out the article “Group VR experiences can produce ego attenuation and connectedness comparable to psychedelics” available open access at www.nature.com/articles/s41598-022-12637-z
Green Tara

Green Tara in progress… Tārā is a buddha manifestation embodying the qualities of feminine energy within the universe. She offers warmth, compassion, and relief from suffering. Like a mother for her children, she engenders, nourishes, and smiles at the vitality of creation. In her Green form, Tārā offers protection from the variety of unfortunate circumstances encountered by beings trapped in cyclic existence. She is characterized by wind energy, enabling her to act quickly to ease the suffering of her children.
White Tara Lockdown Meditation
This White Tara was painted as a lockdown meditation during Jan – April 2021.
Isness-Distributed citizen science studies!

Read all about our latest work working with a global network of citizen scientists to investigate the use of VR to elicit distributed mystical-type experiences which are comparable to pyschedelics! https://arxiv.org/abs/2105.07796


